Location: Home >> Detail
TOTAL VIEWS
Crop Breed Genet Genom. 2019;1:e190002. https://doi.org/10.20900/cbgg20190002
1 Ottawa Research and Development Centre, Agriculture and Agri-Food Canada, 960 Carling Avenue, Neatby Building, Ottawa, ON K1A 0C6, Canada
2 New Liskeard Agricultural Research Station (NLARS), University of Guelph, 340 Armstrong St N, New Liskeard, ON P0J 1P0, Canada
* Correspondence: Weikai Yan.
This article belongs to the Virtual Special Issue "Genetic Gains in Plant Breeding"
Data for multiple traits are routinely collected in crop variety trials and agronomic management studies, which are used to identify superior genotypes, managements, or genotype-management combinations for a given crop and region. However, dealing with multiple traits has been a challenge due to unfavourable associations among traits. In this paper we demonstrate the use of GYT (genotype by yield*trait) analysis for evaluating genotypes (or agronomic managements or genotype-management combinations) based on multiple traits. Genotype evaluation using GYT analysis involves several steps. First, convert the genotype by trait two-way table into a GYT table. Second, standardize the GYT table by each yield-trait combination. Third, apply proper weights to the yield-trait combinations according to their relative importance. Finally, calculate the GYT index, which is the mean across weighted yield-trait combinations for each genotype. The GYT index is a measure of the overall superiority and can be used to rank the genotypes. The GYT index is superior to traditional selection index because it is based on the concept that yield is the most important trait and the economic value of a level of other traits increases with the yield level it is combined. Selection based on the GYT index can prevent low-yielding genotypes from being selected and recommended. GYT analysis can be greatly assisted by the use of GYT biplot, which graphically ranks the genotypes on their overall superiority and shows their strengths and weaknesses. Agronomic managements and genotype-management combinations can be evaluated similarly. Multiple trait data from an oat (Avena sativa L.) management study involving three cultivars, four N-fertilizer treatments, and two fungicide treatments, conducted in northern Ontario in 2015 and 2016, were used to demonstrate GYT analysis.
ATC, average-tester-coordination; GT, genotype by trait; GYT, genotype by yield*trait
Breeding and agronomic management studies share the same goal to improve the yield and quality of crops. Analysis of the data from crop variety trials or agronomic management studies has often been limited to single traits (usually yield), while decisions on the choice of genotypes and/or managements must consider multiple traits. Independent culling and index selection are common approaches utilizing multiple traits in genotype evaluation [1,2]. Independent culling is to discard genotypes whose levels are below a minimum requirement for any trait, no matter how good they are in other traits. Index selection is to select superior genotypes based on an index, which is a linear combination of the target traits. In index selection the economic value of the level of a trait is regarded as independent of other traits. In reality, however, the economic value of the level of a trait is dependent on the levels of other traits, particularly that of yield. For example, superior lodging resistance is valuable only when it is combined with high yield, and it has no economic value if it is associated with very low yield. Similarly, a high level of a quality trait is valuable only when it is combined with high yield; a genotype with superior quality but very poor yield will not be accepted as a cultivar. Thus, the economic value of a trait level is higher when it is associated higher yield. In fact, the purpose of plant breeding is to combine high and reliable yield with desirable levels of other traits in the same genotype. Likewise, the purpose of agronomic management is also to achieve both high yield and good quality. Based on this paradigm, a genotype by yield*trait (GYT) biplot approach for genotype evaluation on multiple traits was developed [3]. The GYT biplot graphically ranks genotypes based on their superiority in combining yield with other target traits and at the same time shows the strengths and weaknesses of the genotypes.
The objectives of this paper are to present an updated version of GYT biplot analysis by allowing differential weights for the yield-trait combinations and to extend it to the analysis of multi-trait data from agronomic management studies. A dataset from an intensive management study of covered oat (Avena sativa L.) in northern Ontario involving different genotypes, nitrogen fertilizer levels, and fungicide treatments will be used to demonstrate the methodology.
The sample dataset used in the case study was taken from an intensive oat management study conducted on a clay-loam soil in New Liskeard, Ontario (47°31'9.86" N, 79°40'14.78" W) in 2015 and 2016. The purpose of the study was to identify superior oat cultivars, agronomic managements, and their combinations, for use in milling oat production in northern Ontario. In addition to yield, good lodging resistance, high levels of test weight, groat content, β-glucan content, and protein content and low levels of oil are considered desirable for milling oat [3–5].
The study was a factorial experiment involved four nitrogen levels (N), three oat cultivars, two fungicide treatments, and two growth regulator treatments. A split plot design with three replications was used, with N levels as the main plots and fungicide, growth regulator, and genotype as the first, second, and third-order sub-plots. The total number of yield plots (the smallest experimental units) each year was 216. Three growth regulator treatments were planned but only two levels were actually applied. As a result, the control treatment for the growth regulator factor was replicated six rather than three times. The four N levels were: control (0N), 60 kg•N•ha−1 applied at planting (60N), 90 kg•N•ha−1 applied at planting (90N), and 60 kg•N•ha−1 at planting plus 30 kg•N•ha−1 at the flag leaf stage (60N + 30N). The source of N was Twin Pine Lawn Fertilizer (34-0-0). The N application at planting was performed with a FABRO seed drill by driving a second time over the plot immediately after planting. The N application at the flag leaf stage was top-dressed by hand. The three cultivars were AC Dieter, a popular but phasing-out milling oat cultivar in eastern Canada, CDC Morrison, a short, early cultivar with exceptionally high β-glucan and protein concentrations but relatively low yield and groat content, and CS Camden, a cultivar becoming increasingly popular in western Canada for its relatively high yield potential, good lodging resistance, and moderately high β-glucan content. The two levels of fungicide were with (Fung) and without (0Fung) applying 0.5 L•ha−1 Twinline (BASF Canada Inc, Mississauga, ON, Canada) at the flag leaf stage. The levels of plant growth regulator were with (2GR) and without (0GR) applying Manipulator (Manipulator 62, Taminco US Inc., Allentown, PA, USA) at the flag leaf stage. The plot size was 5.2 m2 (8 rows with 17 cm between rows). The seeding rate was 330 viable seeds m−2. The sowing dates were 14 May and 5 May, and the harvesting dates were 1 Sept. and 26 Aug. in 2015 and 2016, respectively. Weeds were controlled by applying 1.25 L•ha−1 Logic M (IPCO, Winnipeg, MB, Canada). The crop series were alfalfa-alfalfa-soybean-oat in 2015 and soybean-perennial ryegrass-spring wheat-oat in 2016.
Yield, lodging score, and test weight were determined for each plot. Lodging was rated in a 0 to 9 scale, where 0 means no lodging and 9 means lodged to flat. No lodging occurred in 2016. Bulk samples across replicates for each treatment (i.e., each cultivar-nitrogen-fungicide-growth regulator combination) were determined for groat content using a Codema dehuller, and the groat was ground and scanned using a FOSS DS2500 NIR reflectance unit to predict β-glucan, oil, and protein concentrations based on calibrations developed in-house in Ottawa Research and Development Centre, Agriculture and Agri-Food Canada. The R2 and standard error of the predictions were estimated to be 0.98 and 0.37 for protein, 0.95 and 0.20 for oil, and 0.85 and 0.33 for β-glucan, respectively.
Analysis of variance (not presented) indicated that the growth regular treatment did not have significant effects on yield, any of the quality traits, or lodging score and did not interact with genotypes, nitrogen levels, or fungicide treatments. So, growth regular treatments were treated as extra replications for the other three factors. The quality data, although unreplicated relative to each treatment, are abundantly replicated with regard to each genotype, N-level, fungicide level, and their combinations. The mean values for each genotype-nitrogen-fungicide combination for each of the traits, across the replications, growth regulator treatments, and years, are presented in Table 1. This is the sample dataset to be used to demonstrate GYT analysis. There were significant genotype by year interactions for yield, groat, β-glucan, and protein and significant nitrogen by year interactions for yield, β-glucan, and oil content (results not presented). These interactions indicate that multiple-year experiments are required to make a conclusive decision regarding the treatments. Therefore, it is justified to summarize the data across years.
A genotype by trait (GT) biplot is a graphical presentation of a GT table (Table 2). The method for generating and interpreting a GT biplot was described in [3,6].
Genotype by Yield*Trait (GYT) BiplotA GYT biplot was generated the same way as a GT biplot except that the term “trait” was replaced by the term “yield*trait” (used interchangeably with the term “yield-trait combination”). GYT biplot analysis includes the following four straightforward steps.
Step 1: convert the GT table (Table 2) to a raw GYT table (Table 3). In the GYT table each column is a yield-trait combination. For traits that were so measured that a larger value means more desirable, the yield-trait combination was denoted as “Y*Trait”. For example, the “Y*TW” column contains values from multiplying the yield value (Y) with the test weight value (TW) for each genotype. For traits that were so measured that a larger value means less desirable, the yield-trait combination is denoted as “Y*Trait(-)”, e.g., Y*Lodging(-). The Y*Lodging(-) values were calculated as follows. The lodging score was first transformed to “lodging resistance” such that all genotypes had non-negative values and a larger score means better lodging resistance. Specifically, the transformation was conducted by: Xi = Max + Min − xi, where xi and Xi are the original lodging score and the transformed lodging resistance for genotype i, respectively, and Max and Min are the maximum and minimum lodging scores, respectively. The transformed values are then multiplied with the yield value to obtain the “Y*Lodging(-)” value for each genotype. There may be traits that have an optimal value. For example, although the milling oat industry requires oat oil concentration to be lower than 8.0%, it is not necessarily the lower the better; an oil concentration of 6.0%, say, may be considered as optimal. So the yield-oil combination was denoted as “Y*Oil(~)” and the transformation was done by Xi = Max − |xi − Opt|, where Opt is the optimal value for the trait. This transformed value was then multiplied to the yield value to form the “Y*Oil(~)” values for each genotype. Micronaire index in cotton is another example trait that has an optima value (round 4.0) [7]. The data transformations guarantee that in the GYT table a larger value always means more desirable. The units in the GYT table are not important since the data will be standardized before genotype evaluation.
Step 2: standardize the raw GYT table (Table 3) to form the standardized GYT table (Table 4). This is done by centering (i.e., subtracting the mean) and dividing the centered value by the standard deviation within the respective yield-trait combinations [3] (Yan and Frégeau-Reid, 2018). A GYT index can be calculated from this standardized GYT table for each genotype, which is the mean across all standardized yield-trait combinations (last column of Table 4).
Step 3: apply weights to each yield-trait combination. The traits, or the yield*trait values, were given equal weights in [3]. This can be justified when decisions are made on the GYT biplot. However, when decisions are made solely on the GYT index, then allowing different weights for different traits may be needed if the traits are of different importance. The weights are set as follows: the default weight for a trait is set to 1.0 (but can be changed to anything between 0 and 2); for a trait considered more important, the weight can be set between 1.0 and 2.0; for a trait considered less important, the weight can be set between 0.0 and 1.0. Different researchers may have a different set of weights for the same traits, of course. For the case study, the following weights were assigned: β-glucan (1.0), groat (1.0), test weight (1.0), lodging resistance (1.5), oil (0.5), and protein (0.5). The interface of the software used in this study for trait selection and weight assignment in GYT analysis looks like Figure 1. Applying these weights to the unweighted GYT table (Table 4) led to a weighted GYT table in Table 5.
Step 4: display the weighted (or unweighted) GYT table in a GYT biplot for visual analysis as described in [3].
Evaluation of Different Management Levels Based on Multiple TraitsAgronomic managements can be evaluated using GYT biplot analysis the same way as for genotype evaluation described above. The only change is to replace “genotype” with the treatment factor (e.g., “Nitrogen levels”) or factor combinations (e.g., “Nitrogen and fungicide treatment combinations”).
Evaluation of Genotype by Management Combinations Based on Multiple TraitsGenotype by management combinations can be evaluated using GYT biplot analysis the same way as for genotype evaluation as described above. The only difference is to replace “genotype” with “genotype by management combinations”.
Presented in Figure 2 is the “which-won-what” view of the GT biplot, which graphically displays the GT table (Table 2), after data standardization (as indicated by “Scaling = 1 and Centering = 2” in the upper-left corner of the biplot). The biplot was generated using the GGEbiplot software, in which “Scaling = 1” means the two-way table was scaled by the column (here trait) standard deviation, and “Centering = 2” means the two-way table was centered by column mean [8]. This biplot explained 100% of the variation in Table 2, as indicated at the top-left corner of the biplot, because only three genotypes were involved. It shows that Dieter was best in groat content but poorest in lodging, and it was low in β-glucan and protein concentrations, relative to the other two cultivars; Camden was the highest in yield but poorest in test weight and oil concentration (higher oil is less desirable for milling oat). Morrison was highest in protein content and β-glucan content but lowest in yield and groat content. Thus, the GT biplot effectively reveals the strengths and weaknesses of each genotype. However, it cannot explicitly tell which genotype is the best and should be recommended to oat growers.
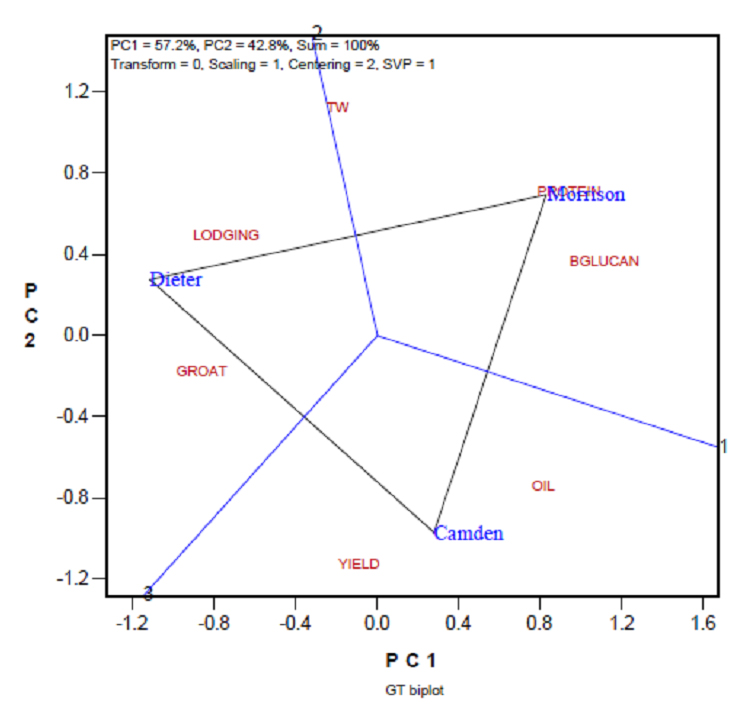 Figure 2. The GT (genotype by trait) biplot to graphically display the genotype by trait table (Table 2). BGLUCAN: β-glucan; TW: test weight.
Figure 2. The GT (genotype by trait) biplot to graphically display the genotype by trait table (Table 2). BGLUCAN: β-glucan; TW: test weight.
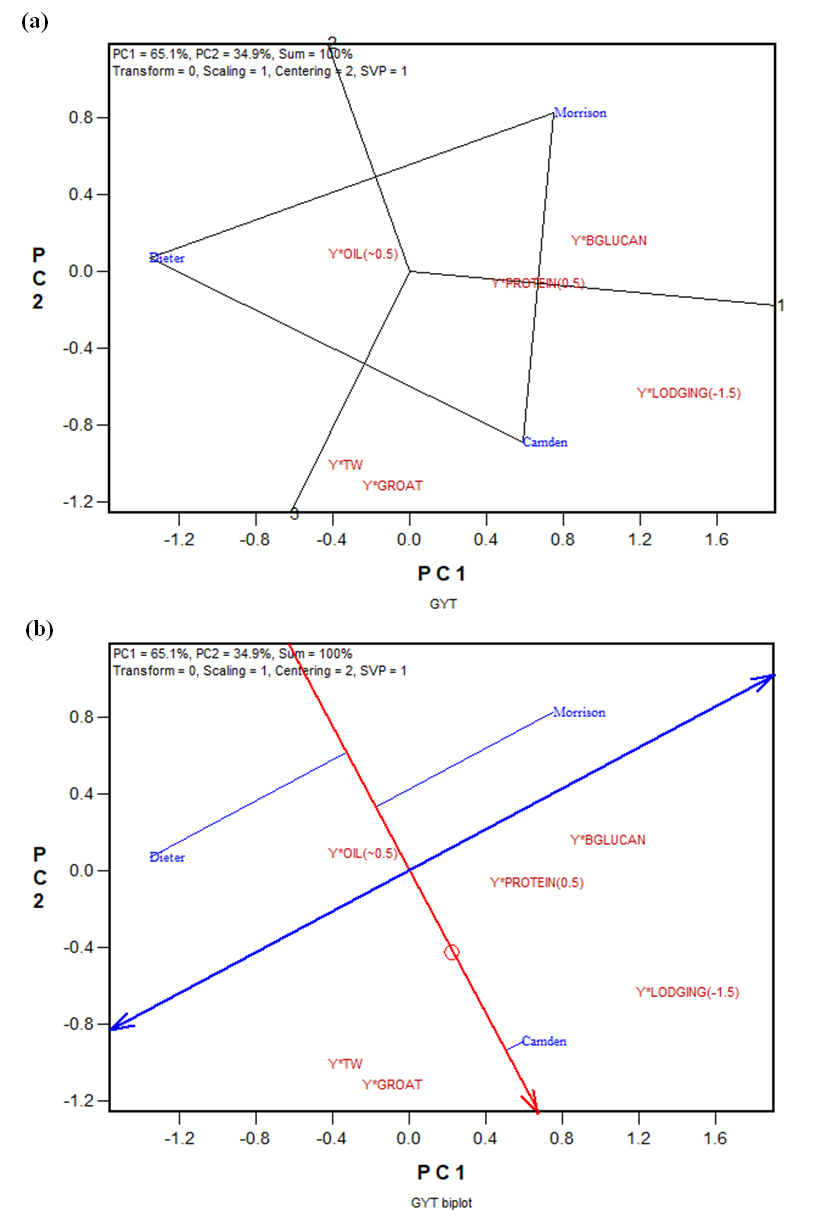 Figure 3. The GYT (genotype by yield*trait) biplot to graphically display the weighted GYT table (Table 5): (a) the which-won-where form of the GYT biplot; (b) the average tester coordination (ATC) form of the GYT biplot. BGLUCAN: β-glucan; TW: test weight.
Figure 3. The GYT (genotype by yield*trait) biplot to graphically display the weighted GYT table (Table 5): (a) the which-won-where form of the GYT biplot; (b) the average tester coordination (ATC) form of the GYT biplot. BGLUCAN: β-glucan; TW: test weight.
Presented in Figure 3a is the which-won-what form of the GYT biplot that displays the GYT table (Table 5). It shows that Camden was the best for all yield-trait combinations except for yield*β-glucan and yield*oil(~0.5). For Yield*β-glucan Morrison was the best and for yield*oil(~0.5) Dieter was the best. Morrison and Camden were similar in yield*protein(0.5). Thus, it is not difficult to conclude that Camden is the best of the three cultivars when all traits are considered. Presented in Figure 3b is the “average-tester-coordination” (ATC) view of the same GYT biplot. The small circle represents the average placement of the yield-trait combinations and is refereed to as the “average tester”. The line with a single arrow passes through the biplot origin and the average tester and is referred to as the “average tester axis” (ATA). The ATA points to higher levels of combination between yield and various traits. The projections of the genotypes onto the ATA are closely correlated with the GYT index (Table 5). Thus, the ATA ranks the genotypes based on their superiority in combining yield with other traits. Camden ranked the best, far better than Morrison and Dieter, and Dieter ranked the poorest (Table 5, Figure 3b). The GYT biplot not only ranks the genotypes on their superiority, but also shows the strengths and weaknesses of the genotypes. For example, it shows that Camden, though overall the best, was weak in yield*oil(~). Although the same conclusions from examining the GYT biplot (Figure 3) can also be arrived by examining the GYT table (Table 5), the GYT biplot is much more effective, particularly when there are many genotypes to compare.
Evaluation of the Nitrogen-Fertilizer Levels Using GT Biplot versus GYT BiplotThe biplot that displays the nitrogen treatment by trait table (not shown) is presented in Figure 4. It shows that the control (0N) had the highest test weight (TW) and lowest lodging score, which are desirable. However, it had the lowest grain yield, groat content, β-glucan concentration, and protein concentration, and the highest oil concentration, which are all undesirable. In contrast, N applications had positive effects including increased yield, groat, β-glucan, and protein and reduced oil, but adverse effects including reduced test weight and increased lodging. These results are consistent with earlier N-fertilizer studies on oat [5,9,10]. So, it is important to find a N treatment or management package so as to maximize the positive effects and minimize the negative effects of nitrogen fertilizer.
In Figure 4, the line perpendicular to the polygon side that connects 90N and 60N + 30N serves the purpose to compare the two N treatments for various traits. It can be seen that although 90N led to slightly higher yield, 60N + 30N led to better levels for everything else, including higher test weight and lower lodging. Thus, split application of the same amount of N (90 kg•ha−1) was better than the one-time application. This conclusion can be better appreciated from the GYT biplot (Figure 5), which graphically displays the weighted nitrogen by yield*trait table (Table 6). Figure 5a shows that 60N + 30N was roughly the same as 90N in combining yield with various traits except lodging resistance. For lodging resistance 60N + 30N was clearly better, although 0N was still the best. The ATC view of the same GYT biplot (Figure 5b) clearly ranks 60N + 30N as the best N treatment, considering all yield-trait combinations. The four N treatments were ranked as 60N + 30N > 90N > 60N > 0N. The main drawback of high level of N (90N) was a poor combination between yield and lodging resistance, which was dramatically alleviated by split N application (Figure 5b). Positive effects of split N application were previously reported for spring wheat (Triticum aestivum L.) [11] and canola (Brassica napus L.) [12,13].
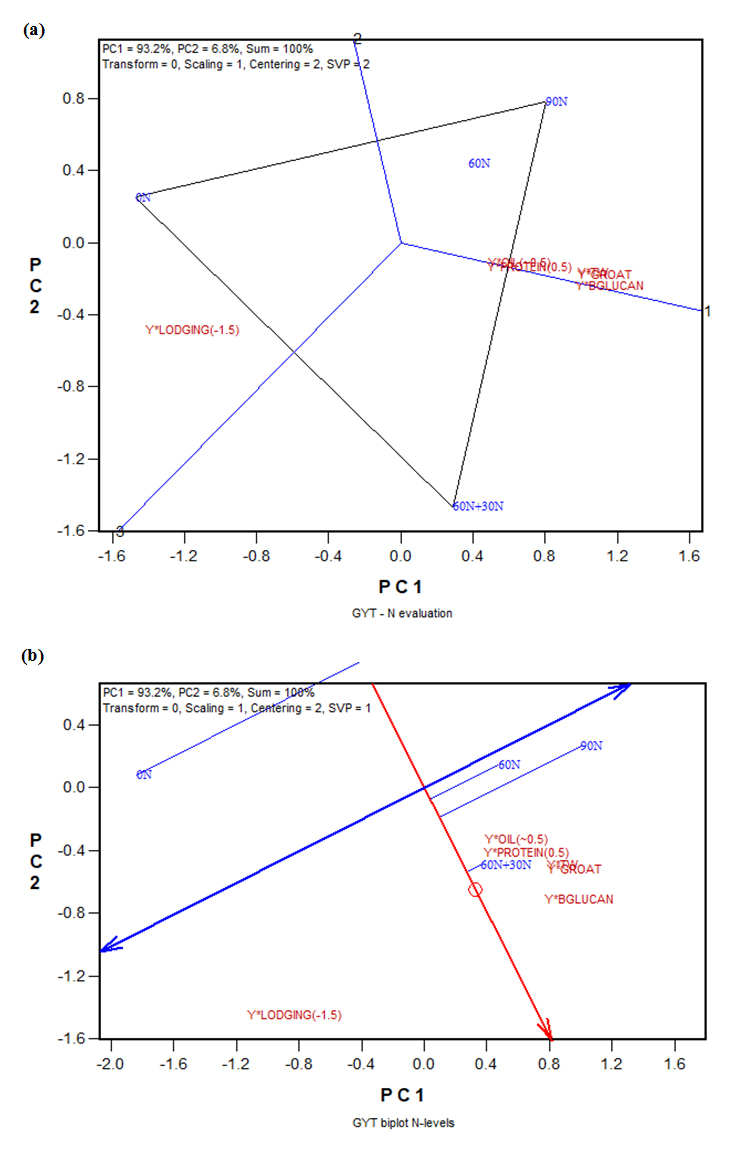 Figure 5. The GYT (genotype by yield*trait) biplot to display the weighted nitrogen by yield*trait (Table 6): (a) the which-won-where form of the GYT biplot; (b) the average tester coordination (ATC) form of the GYT biplot. BGLUCAN: β-glucan; TW: test weight.
Figure 5. The GYT (genotype by yield*trait) biplot to display the weighted nitrogen by yield*trait (Table 6): (a) the which-won-where form of the GYT biplot; (b) the average tester coordination (ATC) form of the GYT biplot. BGLUCAN: β-glucan; TW: test weight.
The nitrogen-fungicide combination by yield*trait biplot presented in Figure 6, based on Table 7, can be used to select the best nitrogen-fungicide combinations on multiple traits. Figure 6a shows that 90N and 60N + 30N, with or without fungicide application, did not differ much in various yield-trait combinations except yield-lodging resistance combination (Y*Lodging(-1.5)). For the latter, 60N + 30N combined with fungicide application (FUNG_60N + 30N) was the best management, as good as 0N, with or without fungicide. Therefore, considering all factors, FUNG_60N + 30N was clearly the best nitrogen-fungicide combination. The second and third best treatment was 90N plus fungicide and 60N plus fungicide (Figure 6b, and Table 7).
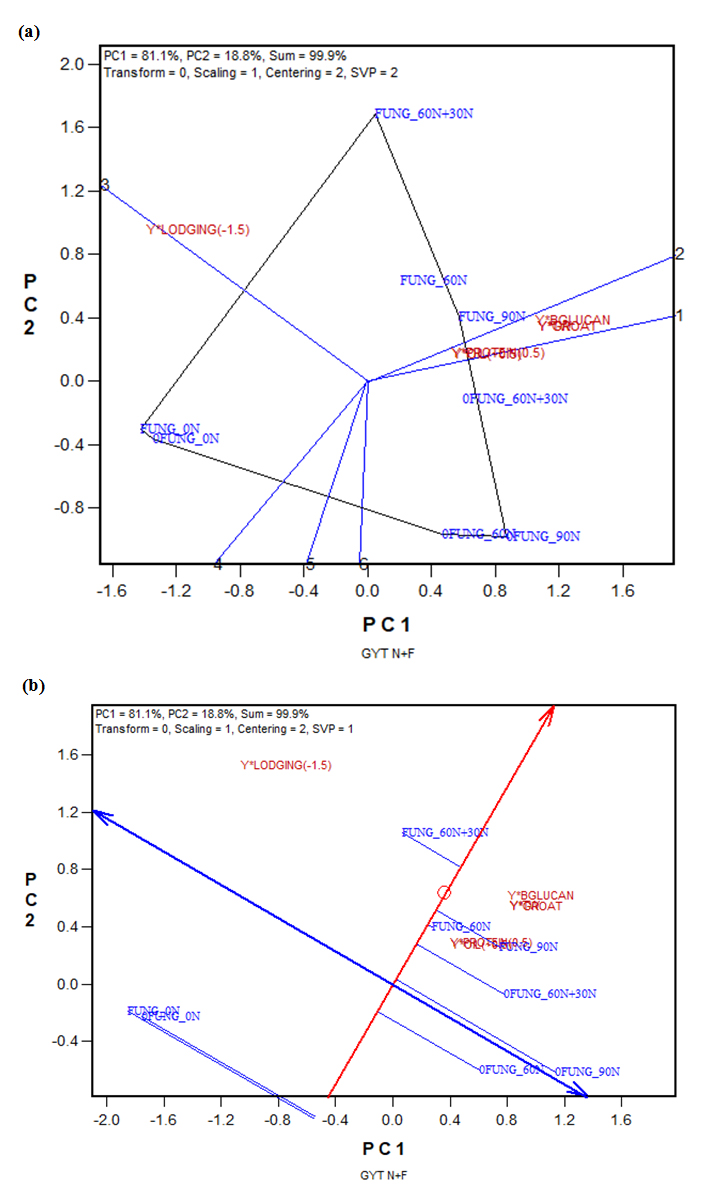 Figure 6. The GYT (genotype by yield*trait) biplot to display the weighted nitrogen-fungicide combination by yield*trait (Table 6): (a) the which-won-where form of the GYT biplot; (b) the average tester coordination (ATC) form of the GYT biplot. BGLUCAN: β-glucan; 0FUNG: without fungicide; FUNG: with fungicide; TW: test weight.
Figure 6. The GYT (genotype by yield*trait) biplot to display the weighted nitrogen-fungicide combination by yield*trait (Table 6): (a) the which-won-where form of the GYT biplot; (b) the average tester coordination (ATC) form of the GYT biplot. BGLUCAN: β-glucan; 0FUNG: without fungicide; FUNG: with fungicide; TW: test weight.
Based on the above analyses, Camden was the best cultivar (Figure 3b) and FUNG_60N + 30N the best management (Figure 6b). Therefore, it can be anticipated that the combination of Camden with FUNG_60N + 30N was the best cultivar-management combination. This is explicitly indicated by the genotype-management combination by yield*trait table (Table 8) and the corresponding GYT biplot (Figure 7b). Figure 7a shows that Camden _60N + 30N_FUNG was the best or close to be the best for all yield*trait combinations except yield*oil.
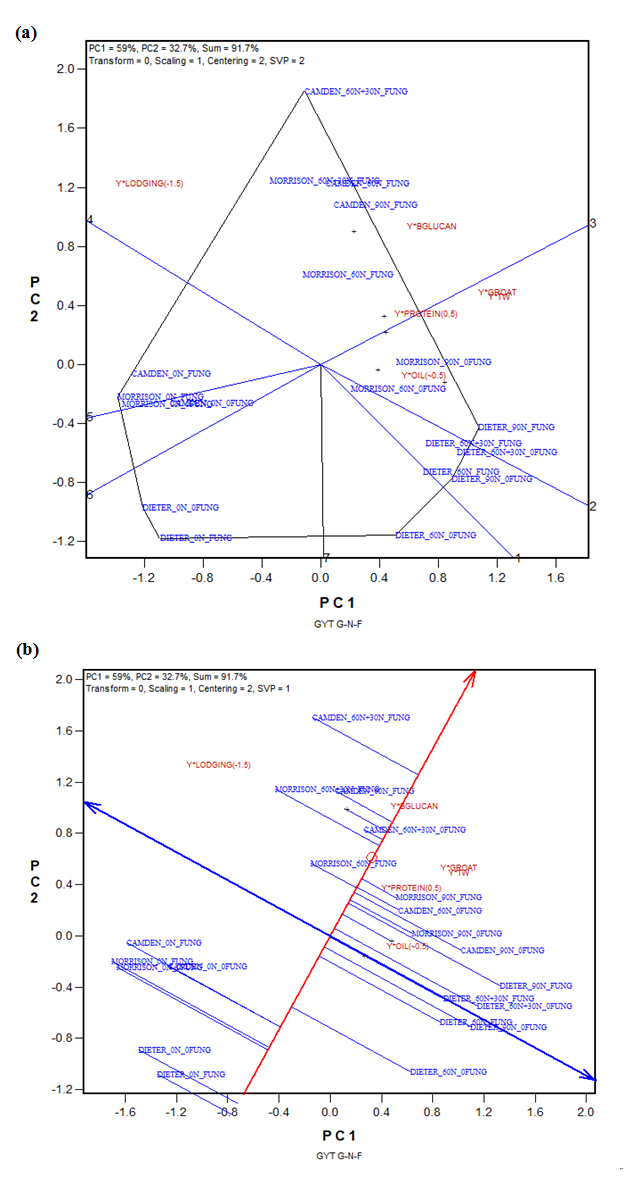 Figure 7. The GYT (genotype by yield*trait) biplot to display the weighted genotype-nitrogen-fungicide combination by yield*trait (Table 6): (a) the which-won-where form of the GYT biplot; (b) the average tester coordination (ATC) form of the GYT biplot. BGLUCAN: β-glucan; 0FUNG: without fungicide; FUNG: with fungicide; TW: test weight.
Figure 7. The GYT (genotype by yield*trait) biplot to display the weighted genotype-nitrogen-fungicide combination by yield*trait (Table 6): (a) the which-won-where form of the GYT biplot; (b) the average tester coordination (ATC) form of the GYT biplot. BGLUCAN: β-glucan; 0FUNG: without fungicide; FUNG: with fungicide; TW: test weight.
Data on multiple traits have always been collected in crop variety trials and agronomic studies. It is also essential to consider all key traits when selecting genotypes and/or agronomic managements. The GYT index is superior to a traditional selection index both conceptually and practically. It reflects the common knowledge and practice that yield is the most important trait; a high level of other traits (e.g., superior lodging resistance or β-glucan content) becomes more valuable when combined with higher yield. The GYT index is calculated as the mean cross all yield-trait combinations (the last column in Tables 5–8), and yield is embedded in every yield-trait combination. Therefore, selection based on GYT index guarantees that selected genotypes (or management or genotype-management combination) are at least high-yielding. In comparison, in a traditional selection index the weight of a trait is fixed and is independent of the levels of other traits; as a result, low yielding genotypes may be selected if they have superior levels in multiple other traits. Such genotypes may be useful as breeding parents but will not be accepted as cultivars. A convenient example is the oat cultivar Morrison included in the case study. Morrison has many good characteristics: exceptionally high β-glucan content, exceptionally high protein content, early maturity, short statue, and good lodging resistance. However, its use as a cultivar has been greatly limited by its low yield potential.
GYT Table versus GYT BiplotSince yield is a component in each yield-trait combination, the various yield-trait combinations tend to be positively correlated or uncorrelated, even though the traits per se are negatively corelated. This feature allows the genotypes (or managements or genotype-management combinations) to be graphically evaluated using the ATC view of the GYT biplot (Figures 3b, 5b, 6b and 7b). Although a GYT table contains all information needed for genotype evaluation, the GYT biplot is clearly more effective. It ranks the genotypes based on their levels of yield-trait combinations (the GYT index), shows the trait profiles, i.e., strengths and weaknesses of the genotypes, and shows the similarities/dissimilarities among the genotypes for various traits. Without the help of the biplot, some important patterns in the table may go unnoticed.
Biplot is a graphical presentation of principle component analysis (PCA) of a two-way table. The use of PCA is to extract “patterns” and get rid of the “noise” in the data. This is why the two-way table estimated from an optimal number of PCs are considered more accurate than the original values in the two-way table [14,15]. From this perspective it is important to determine the number of PCs required to approximate the data. The use of a 2-dimentiaonl biplot is, however, to graphically summarize the patterns of the data, with the awareness that some patterns may not be represented in the biplot. Nevertheless, it is guaranteed that the biplot always displays the most important patterns in the data, which is usually sufficient to make correct decisions. This is because although the data can be infinitely complex, the decision regarding a genotype or treatment must be “yes or no” [8]. GYT analysis can be based either on the GYT table or the GYT biplot. The biplot can be used as a graphical tool to quickly visualize the ranking of the genotypes (or treatments) and their strengths and weaknesses; the table form can be used to verify the conclusions from the biplot. The biplot and the table are usually consistent; in cases when there are large discrepancies, which rarely occurs, the tabulated results should be used for final decisions.
The functionality of the GYT biplot in ranking genotypes (or managements) for their overall superiority is unique; the GT biplot (e.g., Figures 2 and 4) also shows the trait profiles of the genotypes but does not have a meaningful ATC view and cannot be used to rank genotypes or managements.
How Many Traits Should Be Included in GYT Analysis?There is no limit as to the number of traits to be included in the GYT analysis although at least three yield-trait combinations are required for biplot display. It is important to note that the GYT index is dependent on the number of traits that are considered in the analysis. Because yield is included in every yield-trait combination, including more traits in the analysis means a greater weight for yield relative to the other traits, and including fewer traits means a smaller weight for yield. Therefore, the researcher must carefully consider which traits to include in the analysis. Typically, many traits or parameters are determined in crop variety trials and agronomic studies. However, not all traits are equally important. Traits may be classified into three types: I) traits that can be used in positive selection (i.e., in selecting superior genotypes); II) traits that cannot be used in positive selection but can be used in negative selection (i.e., in culling inferior genotypes); and III) traits that may add value but cannot be used in either positive selection or negative selection due to limited importance. For most crops, yield is the only Type-I trait, because a very high-yielding genotype will always be useful and a very low-yielding genotypes will always be unacceptable. Other key target traits are of Type-II. Examples are lodging resistance, test weight, groat content, β-glucan content, and oil content for milling oat. Although a desirable level in any of these traits is not sufficient to make a genotype a superior cultivar, a poor level can limit the adaptation of a cultivar to a target environment or end-use. All other traits may be regarded as of Type-III. It is advised to include only Type-II traits in GYT analysis, in addition to yield (the sole Type I trait). When there are only two traits (in addition to yield) to be considered, then a biplot is not possible, and traditional index selection should be used.
Weighted versus Unweighted GYT AnalysisIn the original paper of GYT biplot analysis [3], the yield-trait combinations were unweighted, i.e., they were given equal weights. This was justified because the focus in that paper was on using all information presented in the GYT biplot, rather than depending on the GYT index alone, for decision making. Because the GYT biplot contains all information about the genotypes on each and every yield-trait combination, it allows proper decision making on what genotypes to select according to a specific target environment and/or end-use. The relative importance of the traits would differ with a different target environment or end-use, and a single set of weights cannot meet all requirements. Nevertheless, for a specified target environment and end-use, and when the focus is on producing a GYT index to rank the genotypes (or managements or genotype-management combinations) based on multiple traits, an option for setting differential weights for different traits and therefore yield-trait combinations will be very useful (Figure 1).
GYT Analysis and Independent CullingFor identifying superior genotypes (or managements or genotype-management combinations), selection using a GYT index and independent culling are both needed to prevent genotypes with serious defects from being selected. The original genotype-trait table (Table 2) can be used for independent culling and the GYT index (Table 5) and the GYT biplot (Figure 3) can be used for index selection. In practice, we rank the genotypes first based on their GYT index, and then discard those that have severe defects for any key traits.
Measure of UncertaintyAs with a traditional selection index, the GYT index does not have a measure of uncertainty as to the rank of the genotypes or managements or genotype-management combinations. However, with some common sense, the graphical presentation of the rank (Figures 3b, 5b, 6b and 7b) can give a fairly good idea on how different two genotypes are in their overall superiority. For example, it should not be difficult to tell that Camden was clearly better than Morrison and Dieter (Figure 3b). This is consistent with the reality that Camden is becoming the number-one oat cultivar in western Canada and is rapidly replacing Dieter as a milling oat cultivar in northern Ontario. Figure 5b shows that split application of the 90 kg•ha−1 N fertilizer (60N + 30N) was clearly better than the one-time application (90N), while the latter did not differ much from 60N. Likewise, Figure 6b shows that the split N application plus the fungicide application (60N + 30N_FUNG) was clearly the best management, and Figure 7b shows that Camden administered with split N application and fungicide application (CAMDEN_60N + 30N_FUNG) was clearly the best genotype-management combination. Analysis of variance can be conducted to see if two genotypes or treatments are statistically different in their GYT index, treating each yield-trait combination as a replication. However, the validity of this test remains to be studied. It is a legitimate question weather the gain from split N application and fungicide application was worthy of the cost due to additional operations; this is out of the scope of this paper, however.
The GYT biplot approach is superior to traditional index selection in genotype evaluation and/or agronomic management evaluation based on multiple traits. First, it is based on the concept that yield is the most important trait and the economic value of the level of all other traits depends on the yield level with which it is associated. As a result, the GYT index from GYT analysis guarantees that selected genotypes and/or managements are superior in combining yield with other target traits and prevents low yielding genotypes and/or managements from being selected and recommended. The GYT biplot not only graphically ranks the superiority of the genotypes and/or managements in combining yield with other traits but also shows the strengths and weaknesses of each genotype and/or management. Therefore, GYT biplot analysis is an effective and informative tool for genotype and management evaluation on multiple traits.
JL and NM designed and conducted the field experiments, JF determined the quality of the oat samples, WY developed the analytical method, conducted the analysis, and drafted the manuscript. All approved the final manuscript.
We declare no conflict of interest in any form.
The data used in the case study was from a study jointly funded by Agriculture and Agri-Food Canada (AAFC), BASF, Canterra Seed, Engage Agro, Grain Farmers of Ontario (GFO), Northern Ontario Farm Innovation Alliance (NOFIA), Quaker Oats/PepsiCo, SeCan, Syngenta, and University of Guelph.
1.
2.
3.
4.
5.
6.
7.
8.
9.
10.
11.
12.
13.
14.
15.
Yan W, Fregeau-Reid J, Mountain N, Kobler J. Genotype and management evaluation based on Genotype by Yield*Trait (GYT) analysis. Crop Breed Genet Genom. 2019;1:e190002. https://doi.org/10.20900/cbgg20190002

Copyright © 2020 Hapres Co., Ltd. Privacy Policy | Terms and Conditions