Location: Home >> Detail
TOTAL VIEWS
Crop Breed Genet Genom. 2019;1:e190011. https://doi.org/10.20900/cbgg20190011
1 UR1052, Génétique et Amélioration des Fruits et Légumes, Institut National de la Recherche Agronomique (INRA), CS60094, 84143 Montfavet, France
2 UR1115, Plantes et Systèmes de culture Horticoles, INRA, CS40509, 84914 Avignon Cedex 9, France
3 Centre opérationnel de Balandran, CTIFL, 751, Chemin de Balandran, 30127 Bellegarde, France
* Correspondence: Rebecca Stevens.
This article belongs to the Virtual Special Issue "Selected Papers from the 17th International EUCARPIA Conference on Genetics and Breeding of Pepper and Aubergine"
Background: We investigated root architecture and plant biomass traits of 25 aubergine genepool accessions representative of 9 Solanum L. species, including S. melongena L., grown at two sites in South-East France in order to quantify the diversity of root phenotypes, identify correlations between traits and determine the influence of the environment on trait stability.
Method: Aubergine seeds were sown in a mixture of sieved compost and vermiculite in 1m high PVC tubes of 10 cm diameter. Roots and aerial parts were harvested after 4–5 weeks growth. Measured root traits included: root depth and root growth rate, maximum apical diameter, minimum apical diameter, the ratio of daughter to mother root apical diameter, and root inter-lateral distance.
Results:
•
•
•
Conclusions: Root architectural traits show diversity within S. melongena and between S. melongena and related species. We confirm that root maximum diameter one of the traits with the highest heritability, i.e., depends on the genotype and is stable in different environments and that root inter-lateral distance is a plastic trait affected by the environment. The study gives some indications on accessions that could be chosen as rootstocks.
Aubergine (Solanum melongena L.) is one of the main fruit vegetables produced in the world, with 52 million tons produced in 2017, of which 94% is produced in Asia (FAOSTAT 2017), where it is a major source of dietary diversification. Most of the diversity kept in worldwide collections therefore originates from this area, as is the case for the INRA Centre for Vegetable Germplasm [1]. For crop improvement, breeders can also use the diversity available in two African cultivated relatives, Solanum aethiopicum L. and Solanum macrocarpon L., or in other wild relatives.
Over the past decades, as for most fruit vegetables, aubergine breeding has focused on yield, different fruits traits such as regularity of shape and colour, brilliance and size, or postharvest conservation, and plant traits such as reduced growth habit for production in greenhouses and decreased hairiness or prickliness. This means that almost half of the plant, i.e., the root system, has been largely neglected in breeding programs, and its unexplored diversity remains available for breeding, even if some root traits have presumably already been co-selected with traits so far targeted by breeders. Vigorous root systems are also needed as rootstocks in grafting [2] but aubergine rootstock research programs have mainly used available tomato rootstocks instead of using the wide diversity available in germplasm related to aubergine, i.e., Solanum L. species belonging to subgenus Leptostemonum [3,4]. Therefore due to the lack of demand from breeders and the difficulty in describing the root system in an easy and repeatable way, germplasm collections have not been evaluated in terms of the phenotypic diversity available underground.
As root system architecture improvement is an increasingly important topic in improving crop resilience to climate change or new pathogens and in reducing inputs, ways of screening collections are becoming increasingly relevant. Differences in the general development of root systems have been identified between aubergine varieties belonging to different ecotypes [5], and a positive correlation was found between root dry weight and stomatal frequency measured on the adaxial leaf surface [6]. Reciprocal grafting experiments showed the equal importance of shoot and root development in determining fruit number, although fruit mean weight is predominantly influenced by root vigour [7]. However, overall, few data are available to select parental material for breeding or research programs.
In order to uncover the role of root phenotypes in environmental stress responses, several studies in different species have detected QTL following root phenotyping [8]. QTL for root architectural traits have been clearly associated with an increase in the capacity to use soil reserves in rice, wheat and maize [9]. In rice, a root-architecture QTL (root angle) allows maintenance of yield under conditions of drought. The underlying gene has been cloned and encodes a protein of unknown function but which modifies the gravitropic response, showing a link between molecular processes and more global architectural traits such as the direction of root growth [10].
Generally, results from independent studies show that certain root architectural traits are involved in environmental stress responses. An example is root length which is correlated with an improvement in drought stress tolerance in several species [9,10]. However other traits are of interest such as root density, root diameter, emergence angle, root hair density and the distribution of biomass between roots and aerial parts [9]. The improvement of root architecture by the alteration of one specific trait is however difficult because some traits show antagonistic effects compared to others, for example, the trait that favours phosphate absorption, which requires many fine shallow roots, and the trait that favours nitrate absorption, which requires longer, deeper roots [11].
The study of root architecture requires a choice of the traits of interest, a phenotyping method and environment and the relevant genetic material. Furthermore, the interaction with the environment will have an impact on the root traits themselves and together these will affect global plant phenotypes such as vigour, yield and disease resistance. The simple traits chosen in this study have been shown to be of interest to evaluate phenotypic diversity in several species [12–14] and are markers for developmental processes including elongation and branching. The significance of these traits is described in Pagès and Kervella [15]: minimum root diameter (referred to as dMin in this study) indicates the potential to make fine roots that maximize soil-root exchange at low cost to the plant whereas maximum root diameter (dMax) shows the potential to produce strong roots which will grow quickly and explore the soil region. Root inter-lateral distance (ILD) gives an indication of the density of the colonization. We have used these traits to phenotype 25 aubergine accessions including cultivated aubergines and related accessions for a total of 9 different species at two different sites.
Twenty five aubergine accessions were selected from the germplasm kept at the INRA Centre for Vegetable Germplasm, Avignon, France (Table 1). Sixteen accessions of Solanum melongena, were chosen to represent the diversity of geographic and climatic origins together with aerial phenotypic diversity, so as to maximize the chance of observing the root phenotypic diversity available in this cultivated species. The other eight species were taken from within the related germplasm, and include the wild form of Asian eggplant (Solanum insanum L.), its closest African wild relative (Solanum incanum L.) and species currently used as aubergine rootstocks (Solanum aethiopicum Aculeatum Group in Japan, Solanum torvum Sw. in many places or occasionally Solanum sisymbriifolium Lam.). The collection was completed with species shown to have grafting affinity with aubergine (S. aethiopicum Gilo Group, Solanum linnaeanum Hepper & P.-M.L. Jaeger, Solanum macrocarpon and Solanum viarum Dunal) according to results obtained in previous trials carried out in France [16].
Plant CulturePVC tubes of 1 m in height, 10 cm diameter were filled with a mix of 50% (v/v) sieved potting compost (K Substrates Select H 70, Klasmann-Deilmann GmbH, Brème, Germany), 50% (v/v) vermiculite (Vermex, Soprema, France). Each tube was fixed to a 20 cm-square flat support, enabling them to be freestanding. Tubes contained the same dry weight of substrate which was not manually packed down in order to remain well aerated. The substrate level was re-adjusted following addition of 2–3 litres of water to the tube.
Apart from the S.melongena accessions, seeds were soaked for 24 h in a 500 ppm solution of gibberellic acid before sowing. Seeds were sown directly into the substrate in the tubes and following germination, one seedling was retained per tube. Watering with standard nutrient solution (N = 4, P = 2, K = 6, Mg = 2 of EC 1.2 mS/cm) at Site G or water at Site C was adjusted to maintain 30% drainage (approximately 300 mL per day). Germination dates were recorded to enable the age of the plant in days to be calculated on harvesting.
For each accession, 4 plants were analysed at each site. The experiment was carried out at INRA GAFL, Montfavet (site “G”, seeds sown on the 26th April or the 4th May 2018—2 plants per accession for each sowing date) in a glasshouse or at the Centre Technique Interprofessionnel des Fruits et Légumes (CTIFL) at Balandran (site “C”, seeds sown on the 7th or 14th May 2018—2 plants per accession for each sowing date) in a plastic tunnel. Plants and roots at site G were harvested between the 5th and 13th June (for plants sown on 26th April) or the 14th and 19th June (plants sown on 4th May). Plants and their roots at site C were harvested on the 27th June (plants sown on 7th May) or 6th July (plants sown on 14th May). The culture at site C did not include the S. torvum accession because the seeds did not germinate. The environmental characteristics at each site are presented in Table 2 and include air temperature, substrate temperature, relative humidity, photosynthetically active radiation (PAR) and the fertilisation regime. As the environments were not equipped in 2018 for all measurements of environmental parameters, measurements are also included from approximately the same period in 2019.
Root Extraction and SamplingRoots were extracted essentially as previously described [12] after 4–5 weeks once roots appeared at the base of the tube, by submerging the tube underwater and then gently tapping the tube held at a 45° angle so the root “carrot” emerged intact and was collected in a mesh tray just below the surface of the water. Roots were gently washed to remove substrate particles. The aerial part of the plant was separated from the root at the crown. The length of the longest root from crown to root tip was measured. A sample was removed from the roots near the bottom of the tube (large diameter roots) and from the crown of the plant at the tips (fine roots). The root samples were kept in shallow dishes in water at 4 °C for up to 10 days before scanning. Two to three separate roots samples were retained from each plant. The remaining root and shoot structures were weighed before and after being dried at 70 °C for 2–3 days.
Root Scanning and Image AnalysisIndividual roots were carefully spread out in 10 cm square plastic petri dishes containing a thin layer of water and covered with a transparent film. The root was then scanned at 2400 dpi using an Epson Perfection 850 Pro flatbed scanner (Long Beach, CA, USA) in transparent mode. The root sample harvested from the distal part of the root system was used for measurements of maximum diameter (dMax; 5–15 roots measured per scan), inter-lateral distance (ILD, 15–30 distances measured per scan) and lateral diameter (the diameter of the “daughter” root that emerges from the largest root on which dMax is measured, dLat; 15–30 roots measured per scan) using ImageJ software [17]. The sample of fine roots was used for measuring the minimum diameter (dMin: 15–30 roots measured per scan). All diameters were apical diameters, measured near the tip where the root is cylindrical. From these measurements the traits listed in Table 3 were calculated. The number of days of growth from germination to harvest was used to obtain “per day” trait values. The dominance coefficient dLat/dMax is calculated from the average ratio dLat-dMin/dMax-dMin. dLat and dMax are measurements taken from the same scan, dMin is the value obtained from the fine roots of the same plant.
Statistical AnalysisStatistical analysis was carried out using XLStat version 16.02 (Addinsoft, www.xlstat.com) using the Anova function with a Tukey post-hoc test at a 5% significance level for multiple comparison of means. Pearson correlations were calculated based on means calculated from the four plants per genotype at each site. Graphics were produced using the ggplot2 package in R (H. Wickham. ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag, New York, NY, USA, 2016). Broad sense heritability of each trait was calculated using a two-way analysis of variance with the genotype and the site as variables as: H2 = σgenotype2/σtotal2, with σtotal2 = σgenotype2 + σsite2 + σinteraction2 + σresidual2.
The complete set of results of the root architectural traits and fresh and dry weight traits and their derivatives (see Table 3) and statistical analysis are presented in Supplementary Table S1. Our first question concerned the range of variability for the measured phenotypes and their derivatives. We examined the variability within the S. melongena species and the inter-species variability. To simplify this task—and as there was an effect of the site of the majority of the traits, we have focused on the data obtained at one of the two sites, site G, for this part of the results section, the following results sections treat data from both sites. The root traits showing the most significant differences between accessions were dMax, ILD and the derived trait dLat/dMax (Supplementary Table S1). Figure 1 presents the results for dMax (Figure 1A) and ILD (Figure 1B) for the 25 accessions from each of the 9 groups. For the maximum diameter trait, the extreme accessions are MM01602 (S. viarum) which has a maximum root diameter of less than 0.75 mm and an S. melongena accession MM00108bis, a French landrace, which has a maximum root diameter of about 1.3 mm. Interestingly, the variability found in the S. melongena group (16 accessions) is almost as large as the variability found when the 9 wild accessions are included. The results for the Inter-Lateral Distance (ILD) also show that the variability found within the S. melongena group is as large as the variability within the whole dataset. The extremes are MM00163 or MM00108bis (which had the largest dMax) with ILDs of just above 2mm and the accession MM00738 (the genotype E8) which has an ILD of about 5 mm. Figure 1C presents one of the biomass traits: root fresh weight has been chosen because of the correlations revealed in the subsequent results section. For this trait, the wild species S. viarum, S. insanum, S. incanum, S. linnaeanum and S. torvum presented the lowest total root fresh weights after 4–5 weeks growth. However the African cultivated species S. aethiopicum, with S. sisymbriifolium, commonly used as rootstocks, had one of the highest root weights.
The comparison of the accessions grown at each site allowed us to estimate the traits that were less sensitive to environmental conditions, and therefore presumably under greater genetic control, from those that were affected by the differences between the two sites. The statistical analysis for the effect of the genotype, the site and the interaction genotype × site for each of the 19 traits is shown in Supplementary Table S2. Most of the traits are significantly affected by the growing environment (site) except for root length (but not root length/day), dMax and shoot dry weight per day. Furthermore for most traits (exceptions being dLat and the ratio of shoot to root fresh weight) there is an interaction between the genotype and the site.
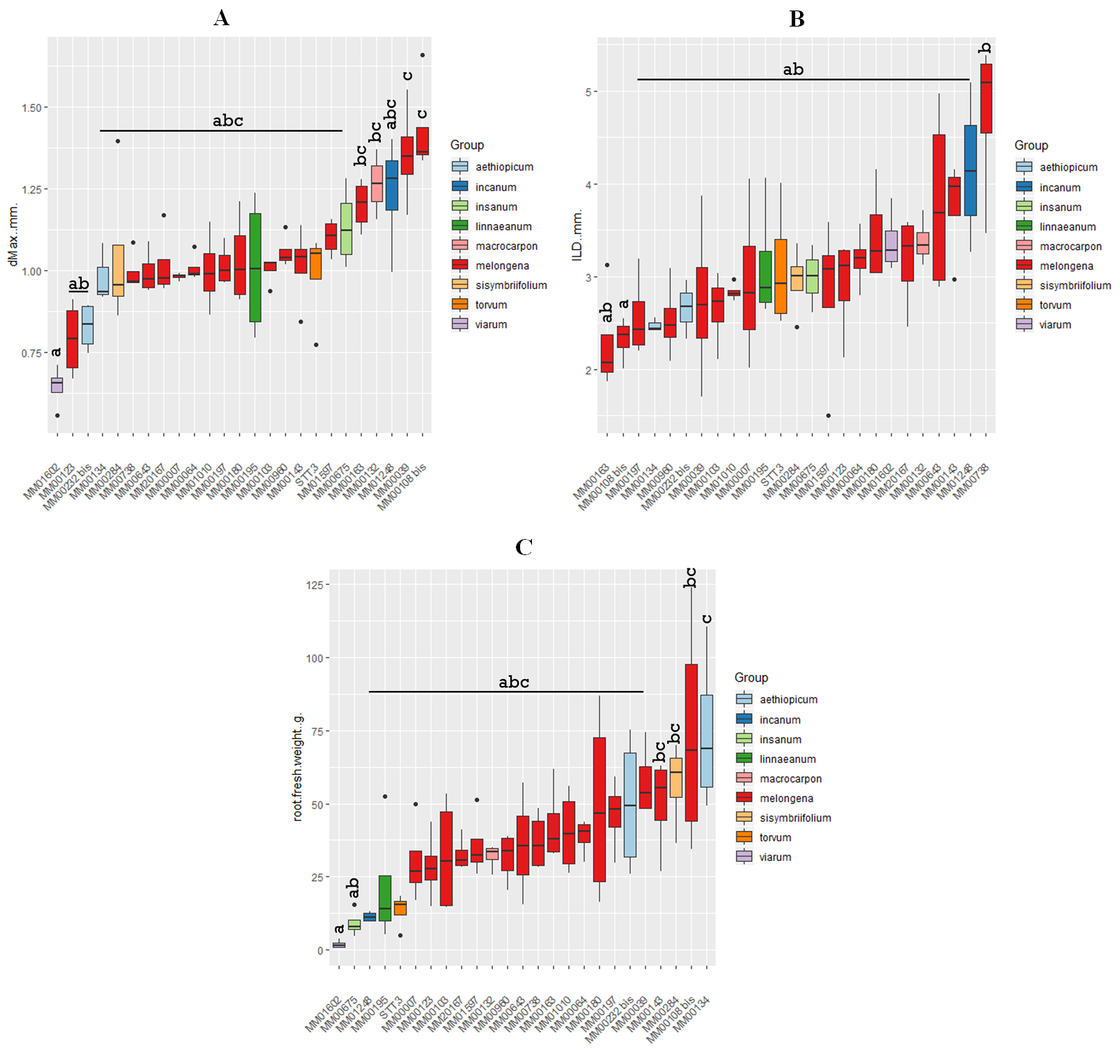 Figure 1. Boxplots representing measurements of maximum root diameter in mm (A), inter-lateral distance in mm (B) and root fresh weight in grams (C). The median is represented by the line across the middle of the box, the end of the box shows the upper and lower quartiles and the vertical line shows the highest and lowest values except for the outliers which are represented by a dot. For each accession a total of 4 independent plants were used. For the maximum diameter and inter-lateral distance, a minimum of two roots were excised from the base of the tube where the thickest roots were found, roots were washed and scanned and a minimum of 15 measurements were made per trait and per scan. For the root fresh weight, roots from four different plants per accession were washed and patted dry before being weighed. The different colours are used to differentiate the nine different species of aubergine and wild relatives. Data presented is from the experiment carried out at Site G. Statistical groups (A, B or C) were assigned following an Analysis of Variance with a post-hoc Tukey test at a 5% significance level.
Figure 1. Boxplots representing measurements of maximum root diameter in mm (A), inter-lateral distance in mm (B) and root fresh weight in grams (C). The median is represented by the line across the middle of the box, the end of the box shows the upper and lower quartiles and the vertical line shows the highest and lowest values except for the outliers which are represented by a dot. For each accession a total of 4 independent plants were used. For the maximum diameter and inter-lateral distance, a minimum of two roots were excised from the base of the tube where the thickest roots were found, roots were washed and scanned and a minimum of 15 measurements were made per trait and per scan. For the root fresh weight, roots from four different plants per accession were washed and patted dry before being weighed. The different colours are used to differentiate the nine different species of aubergine and wild relatives. Data presented is from the experiment carried out at Site G. Statistical groups (A, B or C) were assigned following an Analysis of Variance with a post-hoc Tukey test at a 5% significance level.
The correlations between sites for each trait are shown in Table 4 and lead to similar conclusions. Firstly, not many traits are correlated between the two sites because of the interaction between genotype and site mentioned above: however the traits that do have an R2 above 0.4 and a significant p value are: stem length, shoot fresh weight, root fresh weight per day, shoot fresh weight per day, shoot dry weight per day and total dry weight accumulation per day. Therefore some of the fresh weight/dry weight traits, particularly related to the stem show correlations between the two sites. An analysis of heritability using the data from both sites revealed that dMax and the plant traits (stem length, shoot dry weight and total dry weight accumulation per day) had the highest heritability and were therefore under the most genetic control compared to the other traits.
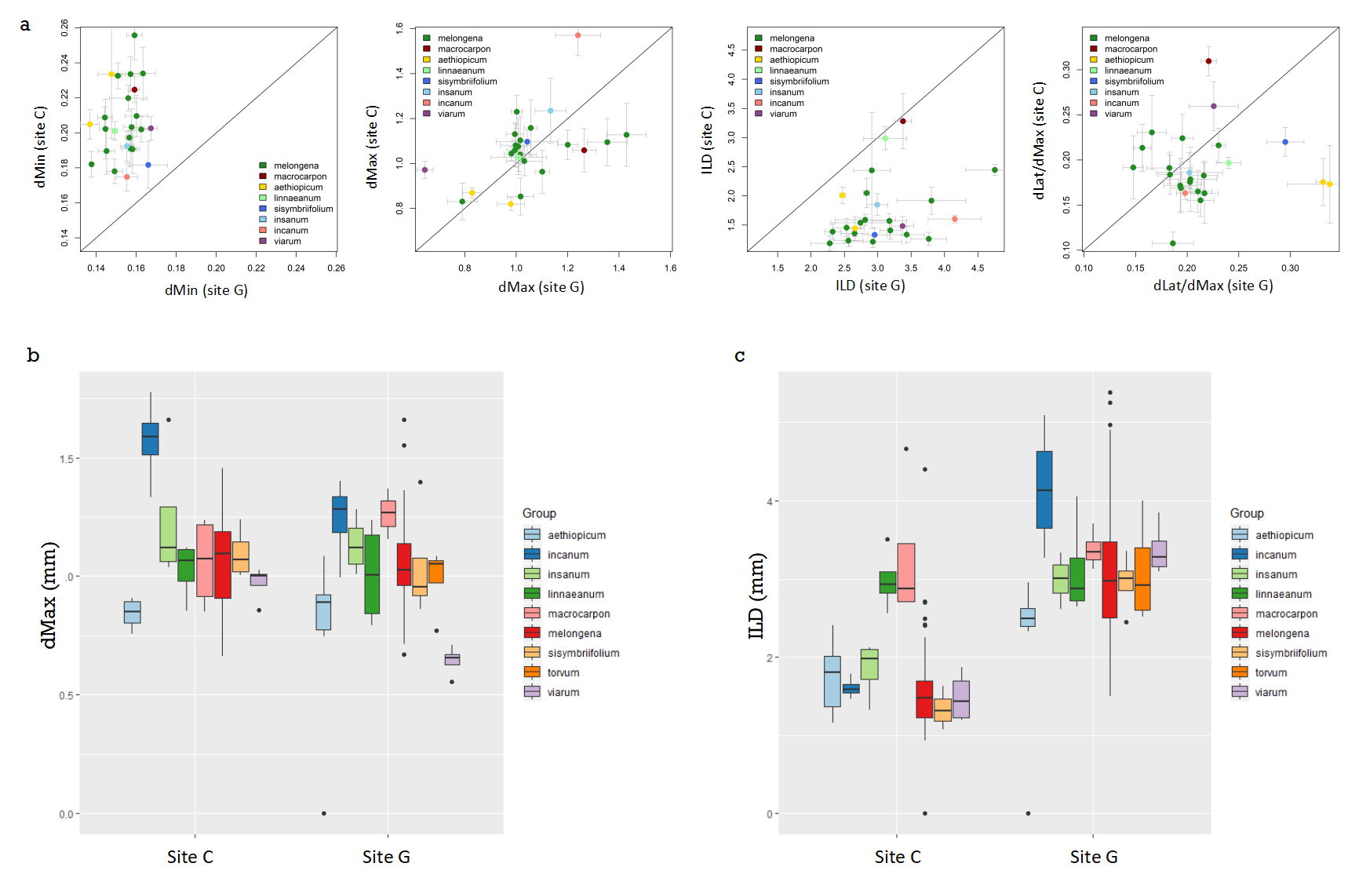 Figure 2. Scatterplots showing correlations between the key root traits (dMin, dMax, dLat/dMax and ILD) at the two sites of culture for each species (a). Boxplots representing measurements of maximum root diameter in mm (b) and inter-lateral distance in mm (c) from experiments carried out at site G or site C. For 2b and 2c, the median is represented by the line across the middle of the box, the end of the box shows the upper and lower quartiles and the vertical line shows the highest and lowest values except for the outliers which are represented by a dot. For the maximum diameter and inter-lateral distance, a minimum of two roots were excised from the base of the tube where the thickest roots were found, roots were washed and scanned and a minimum of 15 measurements were made per trait and per scan. For each accession a total of 4 independent plants were used, results were grouped by species (according to the classification found in Table 1). The different colours are used to differentiate the nine different species of aubergine and wild relatives.
Figure 2. Scatterplots showing correlations between the key root traits (dMin, dMax, dLat/dMax and ILD) at the two sites of culture for each species (a). Boxplots representing measurements of maximum root diameter in mm (b) and inter-lateral distance in mm (c) from experiments carried out at site G or site C. For 2b and 2c, the median is represented by the line across the middle of the box, the end of the box shows the upper and lower quartiles and the vertical line shows the highest and lowest values except for the outliers which are represented by a dot. For the maximum diameter and inter-lateral distance, a minimum of two roots were excised from the base of the tube where the thickest roots were found, roots were washed and scanned and a minimum of 15 measurements were made per trait and per scan. For each accession a total of 4 independent plants were used, results were grouped by species (according to the classification found in Table 1). The different colours are used to differentiate the nine different species of aubergine and wild relatives.
Having established that there is an interaction between genotype and site we can ask whether certain accessions or species show trait stability in the two environments. Figure 2a shows, for the root traits, correlations between the two sites, we note that dMin and ILD are different depending on the site where the measurements were carried out whereas dMax and dLat/dMax appear to be better correlated. Figure 2b shows the detail of the results for dMax for each species, where no significant effect of the site was detected, so the trait distributions for each group are similar: the S. incanum group had the highest values of dMax at each site and the S. aethiopicum group amongst the lowest. In contrast, the ILD trait is affected both by the site and by a genotype x site interaction (Figure 2c; Supplementary Table S2). For this trait, the differences between the groups are not identical for the two sites: it is however notable that the S. linnaeanum and the S. macrocarpon accessions have stable ILD values over the two sites and therefore these accessions show little genotype x site interaction compared to the other groups for this root trait.
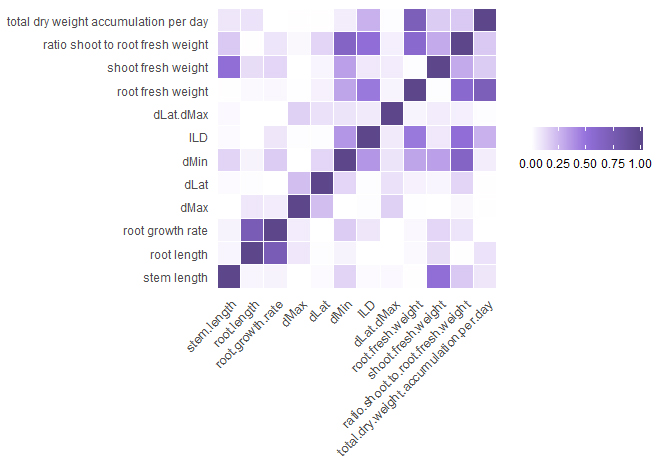 Figure 3. A heatmap showing R2 correlation values for pairs of traits (Table 3) using data from the four plants for each accession grown at each of the two sites (site G and site C; 4 plants per site)
Figure 3. A heatmap showing R2 correlation values for pairs of traits (Table 3) using data from the four plants for each accession grown at each of the two sites (site G and site C; 4 plants per site)
Finally we wished to establish whether correlations existed between roots traits and biomass traits. Figure 3 shows the correlations between the traits of the data combined from the two sites. Many correlations were found between fresh weight and dry weight measurements of the same trait so have not been included in the figure. It is however notable that dMax and dLat and therefore dLat/dMax are not correlated with any other trait whereas for ILD, correlations are found with several traits including root fresh weight. The correlations of dMax and ILD with root fresh weight are shown in Figure 4: the correlation between ILD and root fresh weight is significant (R2 = 0.4685, p = 0) whereas for dMax there is no correlation.
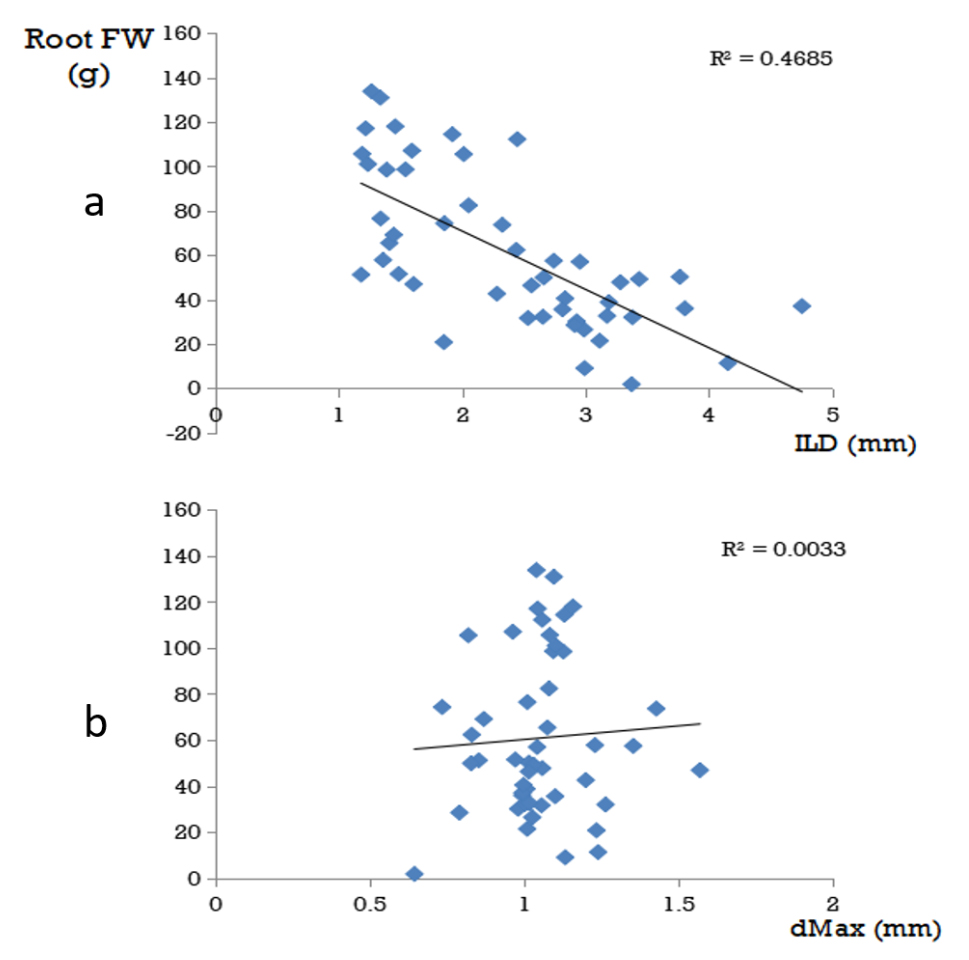 Figure 4. Detail of the correlation shown in Figure 3 for inter-lateral distance (a) and maximum diameter (b) with root fresh weight. Each point on the graph represents the average value of an accession at one of the two sites. The line is the line of best fit for the linear regression and the R2 value for the determination coefficient of the line.
Figure 4. Detail of the correlation shown in Figure 3 for inter-lateral distance (a) and maximum diameter (b) with root fresh weight. Each point on the graph represents the average value of an accession at one of the two sites. The line is the line of best fit for the linear regression and the R2 value for the determination coefficient of the line.
The root and biomass traits chosen for this study show, nearly without exception, variability and sensitivity to the site of culture (environment). Intra-specific variations for the root traits have already been observed in three Solanaceae species including aubergine by Bui et al. [13]. For the current study we purposefully chose to include more accessions of Solanum melongena in order to evaluate the range of variability for this species, being the main cultivated species and more amenable to breeding than some of the related species (which are often difficult to cross or obtain fertile progeny from). Although the S. melongena are over-represented it is still interesting to note that they present a wide range of variability for the root traits compared to the whole panel (Figures 1 and 2), indicating that for these traits domestication has apparently not been too severe in reducing phenotypic variation. It might mean that there is available diversity in S. melongena for breeding for root architectural traits. From this small screening of 16 S. melongena accessions, we have already identified 2 accessions (MM00039 and MM00108 bis) with significantly higher dMax values, which could be used for pre-breeding for this trait.
Although the two sites were similar in terms of climate being only 50 km apart, most traits were affected by the site of culture. Two traits are notable for being stable: these are root maximum diameter, confirming the results of Pagès and Kervella [15], and shoot dry weight accumulation per day which were not statistically affected by the site of culture. These traits also show the highest level of heritability compared to other traits (Table 4). Surprisingly, values for the dMin trait, which has been defined as being a stable trait [15], were larger at site C than site G. One possibility is that the fragile fine roots may have been broken during harvesting. In searching for other reasons to explain the differences in the results at the two sites, we can identify environmental differences: the major one appears to be that at Site G plants were grown in a greenhouse with relatively low PAR light levels compared to site C (Table 2) Photos of the two cultures are shown in Supplementary Figure S1. The compact nature of the plants at site C and the paler green colour of the plants at site G (Supplementary Figure S1) also indicate that plants at site G may have received less light. Certain traits were well correlated between the two sites, these include stem length, shoot fresh weight, root fresh weight per day, shoot fresh weight per day, shoot dry weight per day and total dry weight accumulation per day, again many of these are stem traits: interestingly, stem traits have been identified as hub traits in an analysis of independent studies on herbaceous perennials generating a combined dataset of 23 traits from 2530 individuals of 126 species from 381 plots in Northwest Europe [18], they are also the traits that show high heritability in this paper. Our study also reveals interesting results, such as the observation that the S. linnaeanum and the S. macrocarpon accessions have stable ILD values at the two sites but these observations are based on a limited number of accessions and need to be confirmed in a larger study. The correlations between traits such as the one between ILD and root biomass are in accordance with recent work performed on 10 genotypes of rapeseed [19]: plants that have produced high amounts of biomass are characterized by highly ramified root system architecture. Although these correlation need to be confirmed in a larger study, this may point out diverse strategies among genotypes for soil prospection and resource allocation.
Previous larger studies have included the screening of germplasm for root traits. This is as important as the phenotypic screening carried out for plant and fruit traits related to quality and yield, particularly as root traits are likely to provide tolerance to abiotic and biotic stresses [20,21]. Screening of germplasm has been particularly efficient in crops such as Sorghum where identification of different root system architecture “types” under low and high phosphorus availability can be used to produce varieties potentially adapted to phosphorus scarcity or nitrogen or water limitation [22]. Once phenotyped, germplasm can be used in genetic studies such as genome-wide association if genotyping information is available: an example in pea revealed one significant SNP which was associated with both resistance to A. euteiches and an increased total root projected area, confirming the correlation between resistance and larger root systems in the collection [23]. The confirmation of correlations with agronomic traits also require a larger study than this one although other approaches such as modeling can be used to establish the link between analytical traits and desirable agronomic traits as shown by Pagès and Picon-Cochard [24].
There are numerous phenotyping methods possible for root architecture, [25], each method having specific advantages and disadvantages. The method we have chosen for this study [12] has the advantage of allowing us to work on relatively mature plants (i.e., not seedlings) and so the root architecture has time to develop its potential, we are also able to analyse detailed traits that would not be possible in the field where many roots are broken and fine roots lost. Compared to phenotyping in the field, our method does present certain disadvantages as we do not have an idea of global traits such as the width or volume of the root system. Indeed these are often the traits associated with stress tolerance such as adaptation to drought as illustrated in Comas et al [9], where traits such as increased root biomass, root length density and root depth are associated with drought avoidance in several important crops (rice, maize). It would therefore also be of interest to compare phenotypes obtained in the field for our genotypes.
S. melongena shows a good diversity of root traits and we can identify contrasted traits, certain of which are stable traits (dMax) whereas others are more plastic (ILD). Correlations of ILD with traits related to biomass need to be confirmed in a wider study, compared with measurements of environmental parameters and in different environments such as the field. Interestingly, the African cultivated species S. aethiopicum, with S. sisymbriifolium, had one of the highest root weights in this study which may be a reason for their choice as rootstocks. This study therefore also reveals the S. melongena genotype MM00108bis (LF3-24 Violette de Barbentane) as a potential rootstock candidate, this genotype has one of the highest total root fresh weights and dMax values and one of the lowest ILD values indicating a dense and vigorous root system.
JS and MCD selected and managed the germplasm, JS, VT, CL, AB, CF and RS carried out the experimental work and JS, CL and RS analysed the results, LP designed the phenotyping method and chose the traits. JS, LP, MT and RS obtained funding for the work. RS drafted the manuscript, all authors read and corrected the manuscript.
The authors declare that there is no conflict of interest.
The study was financed by INRA (BAP department—“CARA-COLL” project).
We would like to thank the experiment team at INRA St Maurice for plant management and Cyrille Luquet for tube construction and installation of environmental sensors. We would also like to thank the INRA Centre for Vegetable Germplasm team (CRB-Leg) for providing us with the genetic resources studied here.
1.
2.
3.
4.
5.
6.
7.
8.
9.
10.
11.
12.
13.
14.
15.
16.
17.
18.
19.
20.
21.
22.
23.
24.
25.
Salinier J, Daunay M-C, Talmot V, Lecarpentier C, Pagès L, Bardel A, et al. Root architectural trait diversity in aubergine (Solanum melongena L.) and related species and correlations with plant biomass. Crop Breed Genet Genom. 2019;1:e190011. https://doi.org/10.20900/cbgg20190011

Copyright © 2020 Hapres Co., Ltd. Privacy Policy | Terms and Conditions